Comparative GO Graph (mouse, human, rat, zebrafish)
Compare experimental GO annotations for Human-Mouse-Rat-Zebrafish orthologs to Mouse En2
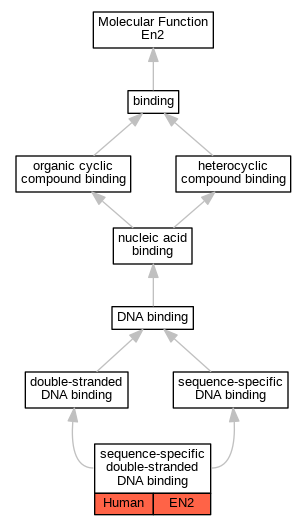
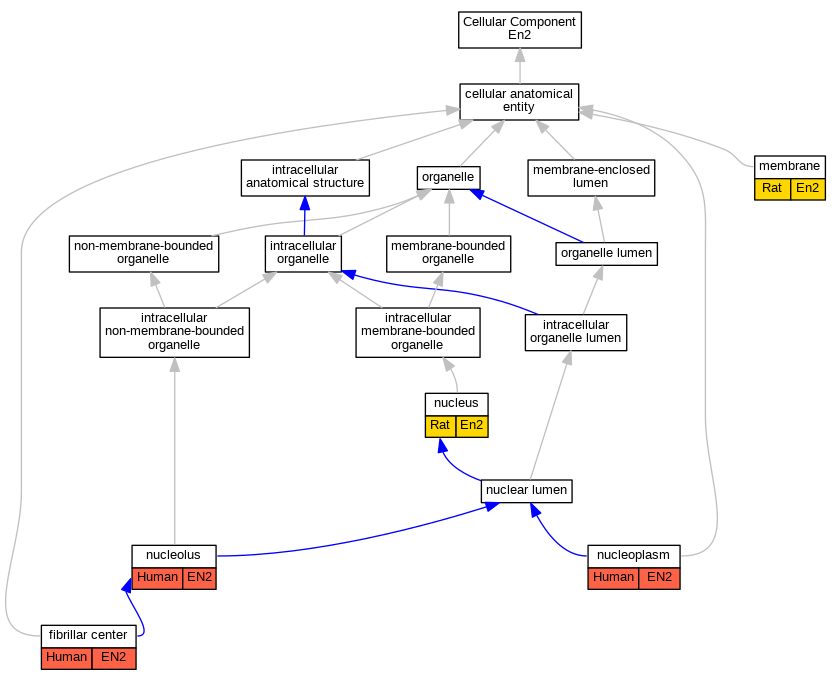
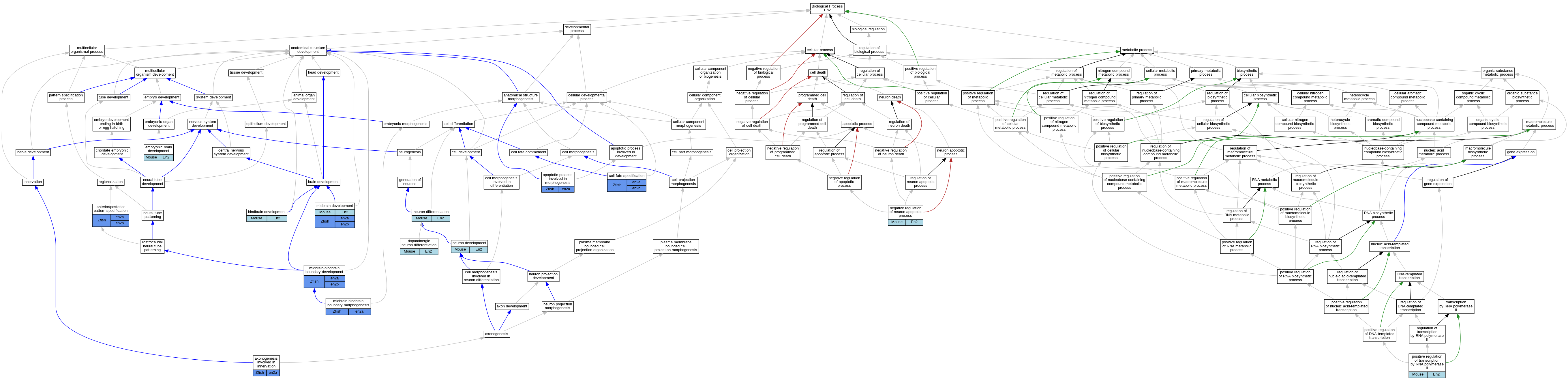
Graphs display curated GO classifications for mouse, human, rat and zebrafish orthologs annotated from the biomedical literature.
- This class contains
- Human genes EN2
- Mouse genes En2
- Rat genes En2
- Zfish genes en2a,en2b
- Annotations are indicated with nodes colored by organism:
Human,
Mouse,
Rat,
Zfish
- Relations between GO terms are indicated by colored edges "is_a"; "part_of"; "regulates"; "positively_regulates"; "negatively_regulates"
- GO annotations: Human from GOA; Mouse from MGI; Rat from RGD; Zebrafish from ZFIN.
- Only experimental annotations are displayed: EXP, IDA, IEP, IGI, IMP, IPI, HTP, HDA, HMP, HGI, HEP. Evidence codes are listed at the bottom of the page.
 Full set of experimental annotations for Human-Rat-Zebrafish genes considered orthologs of Mouse in the Alliance stringent orthology set En2
Full set of experimental annotations for Human-Rat-Zebrafish genes considered orthologs of Mouse in the Alliance stringent orthology set En2
| Category |
ID |
Classification term |
Gene |
Evidence |
Inferred from |
Organism |
Reference |
| Molecular Function | GO:1990837 | sequence-specific double-stranded DNA binding | EN2 | IDA | | Human | PMID:28473536 |
| Cellular Component | GO:0001650 | fibrillar center | EN2 | IDA | | Human | GO_REF:0000052 |
| Cellular Component | GO:0016020 | membrane | En2 | IDA | | Rat | PMID:9169834|RGD:5687201 |
| Cellular Component | GO:0005730 | nucleolus | EN2 | IDA | | Human | GO_REF:0000052 |
| Cellular Component | GO:0005654 | nucleoplasm | EN2 | IDA | | Human | GO_REF:0000052 |
| Cellular Component | GO:0005634 | nucleus | En2 | IDA | | Rat | PMID:9169834|RGD:5687201 |
| Biological Process | GO:0009952 | anterior/posterior pattern specification | en2a | IGI | ZFIN:ZDB-GENE-980526-40,ZFIN:ZDB-GENE-990415-72 | Zfish | PMID:12917294 |
| Biological Process | GO:0009952 | anterior/posterior pattern specification | en2b | IGI | ZFIN:ZDB-GENE-980526-167,ZFIN:ZDB-GENE-990415-72 | Zfish | PMID:12917294 |
| Biological Process | GO:0060561 | apoptotic process involved in morphogenesis | en2a | IGI | ZFIN:ZDB-MRPHLNO-060930-2,ZFIN:ZDB-MRPHLNO-120207-2 | Zfish | PMID:22223680 |
| Biological Process | GO:0060385 | axonogenesis involved in innervation | en2a | IGI | ZFIN:ZDB-MRPHLNO-171117-11,ZFIN:ZDB-MRPHLNO-171117-7,ZFIN:ZDB-MRPHLNO-171117-9 | Zfish | PMID:28807781 |
| Biological Process | GO:0060385 | axonogenesis involved in innervation | en2a | IGI | ZFIN:ZDB-MRPHLNO-171117-10,ZFIN:ZDB-MRPHLNO-171117-11,ZFIN:ZDB-MRPHLNO-171117-6,ZFIN:ZDB-MRPHLNO-171117-7,ZFIN:ZDB-MRPHLNO-171117-8,ZFIN:ZDB-MRPHLNO-171117-9 | Zfish | PMID:28807781 |
| Biological Process | GO:0001708 | cell fate specification | en2a | IGI | ZFIN:ZDB-GENE-980526-40,ZFIN:ZDB-GENE-990415-72 | Zfish | PMID:12917294 |
| Biological Process | GO:0001708 | cell fate specification | en2b | IGI | ZFIN:ZDB-GENE-980526-167,ZFIN:ZDB-GENE-990415-72 | Zfish | PMID:12917294 |
| Biological Process | GO:0071542 | dopaminergic neuron differentiation | En2 | IGI | MGI:MGI:95389 | Mouse | PMID:15175251 |
| Biological Process | GO:0071542 | dopaminergic neuron differentiation | En2 | IGI | MGI:MGI:95389 | Mouse | PMID:15340832 |
| Biological Process | GO:1990403 | embryonic brain development | En2 | IGI | UniProtKB:P09065 | Mouse | MGI:MGI:75252|PMID:7624797 |
| Biological Process | GO:0030902 | hindbrain development | En2 | IGI | MGI:MGI:95389 | Mouse | PMID:15607945 |
| Biological Process | GO:0030901 | midbrain development | En2 | IGI | MGI:MGI:95389 | Mouse | PMID:15175251 |
| Biological Process | GO:0030901 | midbrain development | En2 | IGI | MGI:MGI:95389 | Mouse | PMID:15607945 |
| Biological Process | GO:0030901 | midbrain development | en2a | IGI | ZFIN:ZDB-GENE-980526-40,ZFIN:ZDB-GENE-990415-72 | Zfish | PMID:12917294 |
| Biological Process | GO:0030901 | midbrain development | en2b | IGI | ZFIN:ZDB-GENE-980526-167,ZFIN:ZDB-GENE-990415-72 | Zfish | PMID:12917294 |
| Biological Process | GO:0030917 | midbrain-hindbrain boundary development | en2a | IGI | ZFIN:ZDB-MRPHLNO-060930-2,ZFIN:ZDB-MRPHLNO-060930-3 | Zfish | PMID:11477690 |
| Biological Process | GO:0030917 | midbrain-hindbrain boundary development | en2b | IGI | ZFIN:ZDB-MRPHLNO-060930-2,ZFIN:ZDB-MRPHLNO-060930-3 | Zfish | PMID:11477690 |
| Biological Process | GO:0021555 | midbrain-hindbrain boundary morphogenesis | en2a | IGI | ZFIN:ZDB-GENE-980526-167,ZFIN:ZDB-MRPHLNO-120207-2 | Zfish | PMID:22223680 |
| Biological Process | GO:0043524 | negative regulation of neuron apoptotic process | En2 | IGI | UniProtKB:P09065 | Mouse | MGI:MGI:3578062|PMID:15175251 |
| Biological Process | GO:0048666 | neuron development | En2 | IGI | MGI:MGI:95389 | Mouse | PMID:11312297 |
| Biological Process | GO:0030182 | neuron differentiation | En2 | IGI | MGI:MGI:95389 | Mouse | PMID:15340832 |
| Biological Process | GO:0030182 | neuron differentiation | En2 | IGI | MGI:MGI:95389 | Mouse | PMID:15607945 |
| Biological Process | GO:0045944 | positive regulation of transcription by RNA polymerase II | En2 | IDA | | Mouse | PMID:9102311 |
Gene Ontology Evidence Code Abbreviations:
| EXP Inferred from experiment |
| IDA Inferred from direct assay |
| IEP Inferred from expression pattern |
| IGI Inferred from genetic interaction |
| IMP Inferred from mutant phenotype |
| IPI Inferred from physical interaction |
| HTP Inferred from High Throughput Experiment |
| HDA Inferred from High Throughput Direct Assay |
| HMP Inferred from High Throughput Mutant Phenotype |
| HGI Inferred from High Throughput Genetic Interaction |
| HEP Inferred from High Throughput Expression Pattern |
 Analysis Tools
Analysis Tools





